Simple Skull Stripping Method
Abstract/Motivation
Magnetic resonance images (MRI) play crucial role in radiology and neurology. The high resolution head MRI scans contain some non-brain tissue such as skin, fat, muscles, eyes. The presence of the non-brain tissue hinders performance of automated brain image segmentation and treatment designed methods. The process of removing non-brain tissue from the medical scan is referred as a skull-stripping.
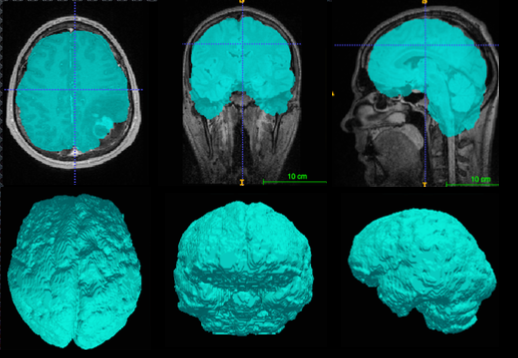 Figure 1: Segmentation of the brain tissue from MRI scan by state-of-the art method (BETs [3]). The presence of a brain tumor reduce method accuracy, leading to under- and/or over-segmentation. Such segmentation artifacts strongly influence performance of other consequent image analysis and treatment planning methods
Figure 1: Segmentation of the brain tissue from MRI scan by state-of-the art method (BETs [3]). The presence of a brain tumor reduce method accuracy, leading to under- and/or over-segmentation. Such segmentation artifacts strongly influence performance of other consequent image analysis and treatment planning methods
 Figure 2: Segmentation of the brain tissue via proposed method. The performance of the method is not compromised by diseases manifestations such as brain tumour, making it a promising method for clinical practice.
Figure 2: Segmentation of the brain tissue via proposed method. The performance of the method is not compromised by diseases manifestations such as brain tumour, making it a promising method for clinical practice.
|
In the past several skull stripping methods were designed. A recent review [1] provides a summary of the existing methods. However these methods are designed either for healthy brains or brains of patients suffering disease that do not significantly alternate the brain structure. The accuracy of these methods drops significantly in the presence of brain lesions and/or resection cavities, which is typical case of the Glioblastoma patients (see Figure 1). Furthermore, the current methods are designed mainly for T1w MRI scans. Thus these methods can not be used in the absence of T1w images, i.e. cases when only PET scans are acquired.
We proposed a simple automated skull stripping method robust against brain diseases alternations such as resection cavity and/or brain lesions (see Figure 2). Moreover the method can be apply to different imaging modalities including MRI, PET and CT.
Thesis objectives
The aim of the project is to bring the proposed method to clinical practice by:
- Testing the proposed methods on public and private datasets ([2])
- Apply machine learning methods to further improve the performance of the method
- Design of a working pipeline and user interface that can be used by clinicians
- Providing open source code, with a tutorial, for public and clinical use
Requirements
- Good programming skill (C/C++, Python and Unix)
- Software design
- Basic knowledge about image processing
- Desire to Learn
- Independent worker
References
[1] Kalavathi P, Prasath VBS: Methods on skull stripping of MRI head scan imagesA review, Journal of digital imaging, 2016 - Springer
[2] Shattuck DW, Prasad G, Mirza M, Narr KL, Toga AW: Online resource for validation of brain segmentation methods.NeuroImage 45(2):431439, 2009
[3] http://poc.vl-e.nl/distribution/manual/fsl-3.2/bet/index.html
Contact
If you are interested in the project or if you have any questions please contact
|
|
 Figure 1: Segmentation of the brain tissue from MRI scan by state-of-the art method (BETs [3]). The presence of a brain tumor reduce method accuracy, leading to under- and/or over-segmentation. Such segmentation artifacts strongly influence performance of other consequent image analysis and treatment planning methods
Figure 1: Segmentation of the brain tissue from MRI scan by state-of-the art method (BETs [3]). The presence of a brain tumor reduce method accuracy, leading to under- and/or over-segmentation. Such segmentation artifacts strongly influence performance of other consequent image analysis and treatment planning methods
 Figure 2: Segmentation of the brain tissue via proposed method. The performance of the method is not compromised by diseases manifestations such as brain tumour, making it a promising method for clinical practice.
Figure 2: Segmentation of the brain tissue via proposed method. The performance of the method is not compromised by diseases manifestations such as brain tumour, making it a promising method for clinical practice.