|
|
Research
My research is mainly focused on the validation, registration, and segmentation of medical images, in particular:- enhancement, segmentation, and registration of angiographic imagery
- vessel modeling
- estimation and analysis of the hemodynamics of blood
- Image-based tracking in fluoroscopic images
- breathing motion compensation in interventional procedures
- analysis, reconstruction, and registration of histology slices
- validation of new imaging technologies via registration to histology
- dynamic analysis of microscopic image sequences
Active research projects
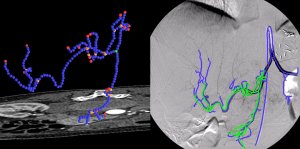
Registration of Angiographic ImagesAngiographic images visualize vascular structure in different modalities like X-Ray, CT, or MR data sets. In many medical applications, a registration and proper visualization of the data sets, especially the vasculature is useful for a better navigation. The focus of this project lies on 2D/3D registration of angiographic data where intensity-based, feature-based, and hybrid approaches are evaluated, the latter two of them requiring an accurate 2D and 3D segmentation of the data. The main clinical partner is the radiology department of the Universitätsklinikum Großhadern (Ludwig-Maximilian Universität München) , industrial partner is Siemens Medical Solutions, Forchheim. |
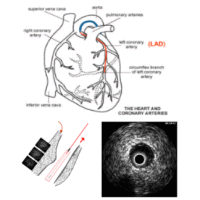
Assessment of Fluid Tissue Interaction Using Multi-Modal Image Fusion for Characterization and Progression of Coronary AtherosclerosisCoronary artery diseases such as atherosclerosis are the leading cause of death in the industrialized world. In this project, we develop computational tools for segmentation and registration problems on intravascular images including IVUS (Intravascular Ultrasound) and OCT (Optical Coherence Tomography). One sample component of this project is Automatic Stent Implant Follow-up from Intravascular OCT Pullbacks. The stents are automatically detected and their distribution is analyzed for monitoring of the stents: their malpositioning and/or tissue growth over stent struts. |
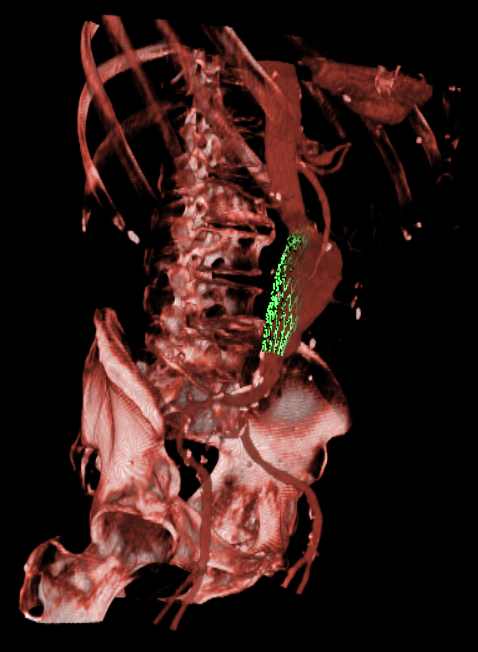
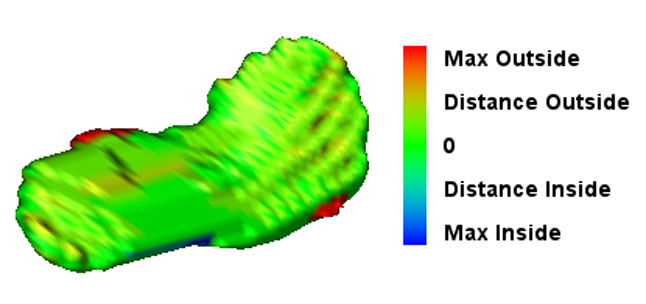
Endovascular Stenting of Aortic AneurysmsEndovascular stenting is a minimally invasive treatment technique for aortic aneurysms or dissections. Thereby, a certain aortic prosthesis (stent graft) is placed inside the aortic aneurysm in order to prevent a life-threatening rupture of the aortic wall. Prior to the intervention, a computed tomography angiography (CTA) is acquired on which the surgical staff can measure the parameter of the desired stent graft and finalize the intervention workflow. The entire interventional catheter navigation is done under 2D angiography imaging where the physician is missing the important 3D information. The purpose of our project is two-fold:1. In the planning phase, a modified graph cuts algorithm automatically segments the aorta and aneurysm, so the surgical staff can choose an appropriate type of stent to match the segmented location, length, and diameter of the aneurysm and aorta. By visualizing the defined stent graft next to the three-dimensionally reconstructed aneurysm, mismeasurements can be detected in an early stage. Our main goal is the creation of an interactive simulation system that predicts the behaviour of the aortic wall and the movement of the implanted stent graft. 2. During implantation of the stent graft, after an intensity based registration of CTA and angiography data, the current navigation can be visualized in the 3D CT data set at any time. This includes solutions for electro-magnetic tracking of catheters as well as guide wires and stent grafts. Eventually, Our main goal is the creation of solutions that enable the surgeon to enhance the accuracy of the navigation and positioning, along with a minimum use of angiography, leading to less radiation exposure and less contrast agent injection. |
Semi-Automatic Patellar Cartilage SegmentationDevelopment and refinement of a software system for semi-automatic segmentation of the patellar cartilage is the main goal of this project. By providing tools for sub-pixel accurate edge tracing, automatic contour completion, and adequate visualization, a remarkable speed-up of the physicians segmentation process can be achieved. Also, improved exactness can be reached for cartilage segmentation if expertise and automation are merged in a meaningful way. |
Assessment of Knee CartilageDegeneration of knee joint cartilage is an important and early indicator of osteoarthritis (OA) which is one of the major socio-economic burdens nowadays. Accurate quantification of the articular cartilage degeneration in an early stage using MR images is a promising approach in diagnosis and therapy for this disease. Particularly, volume and thickness measurement of cartilage tissue has been shown to deliver significant parameters in assessment of pathologies. Here, accurate computer-aided diagnosis tools could improve the clinical routine where image segmentation plays a crucial role. In order to overcome the time-consuming and tedious work of manual segmentation, one tries to automate the segmentation as much as possible. We focus on novel atlas-based segmentation methods for knee cartilage as well as improve todays clinical routine of manual segmentation methods. In addition, we try to evaluate different methods for the assessment of parameters such as volume and thickness which could allow computer-aided diagnosis of knee cartilage pathologies in an early stage. |
Dissertation
Abstract: Angiographic imaging is a widely used monitoring tool for minimally invasive vascular treatment and pathology access. Especially in deforming abdominal areas, the registration of pre- and intraoperative image data is still an unsolved problem, but important in several aspects. In particular, treatment time and radiation exposure to patient and physician can be significantly reduced with the resulting 2D-3D data fusion. The focus of this work is to provide methods for the registration of 2D vascular images acquired by a stationary C-arm to preoperative 3D angiographic Computed Tomography (CT) volumes, in order to improve the workflow of catheterized liver tumor treatments. Fast and robust vessel segmentation techniques are used to prepare the necessary graph data structures for a successful alignment. Here, we introduce restricted correspondence selection and iterative feature space correction to drive the proposed rigid-body algorithms to global and accurate solutions. Moreover, it is shown for the first time that the assignment of natural constraints on vessel structures allows for a successful recovery of a 3D non-rigid transformation despite a single-view scenario. Based on these results, novel volumetric visualization and roadmapping techniques are developed in order to resolve interventional problems of reduced depth perception, blind navigation, and motion blur.| M. Groher 2D-3D Registration of Angiographic Images Technische Universtät München, 2008 (bib) |
Publications
| 2012 | |
| H. Heibel, B. Glocker, M. Groher, M. Pfister, N. Navab
Interventional Tool Tracking using Discrete Optimization IEEE Transactions on Medical Imaging (TMI) (bib) |
|
| Y. Yagi, M. Groher, M. Feuerstein, M. Onozato, H. Heibel, N. Navab
Overcoming Challenges in Histology 3D Imaging 11th European Congress on Telepathology and 5th International Congress on Virtual Microscopy, Venice, Italy, June 2011. (bib) |
|
| A. Chekkoury, P. Khurd, Jie Ni, C. Bahlmann, , Amar Patel, L. Grady, M. Singh, M. Groher, N. Navab, , Jeffrey Johnson, Anna Graham, Ronald Weinstein
Automated Malignancy Detection in Breast Histopathological Images SPIE Medical Imaging, 04-09 February 2012, San Diego, California, USA (bib) |
|
| N. Brieu, N. Navab, J. Serbanovic-Canic, W. Ouwehand
, D. Stemple, A. Cvejic, M. Groher
Image-based Characterization of Thrombus Formation in Time-lapse DIC Microscopy Medical Image Analysis 2012 (bib) |
|
| 2011 | |
| N. Brieu, M. Groher, J. Serbanovic-Canic, A. Cvejic, W. Ouwehand
, N. Navab
Joint Thrombus and Vessel Segmentation Using Dynamic Texture Likelihoods and Shape Prior Medical Image Computing and Computer-Assisted Intervention (MICCAI 2011), Toronto, Canada, September 2011 (bib) |
|
| M. Feuerstein, H. Heibel, J. Gardiazabal, N. Navab, M. Groher
Reconstruction of 3-D Histology Images by Simultaneous Deformable Registration Proceedings of Medical Image Computing and Computer-Assisted Intervention (MICCAI 2011), Toronto, Canada, September 2011. The original publication is available online at www.springerlink.com (bib) |
|
| 2010 | |
| N. Brieu, J. Serbanovic-Canic, A. Cvejic, D. Stemple, W. Ouwehand
, N. Navab, M. Groher
Thrombus Segmentation by Texture Dynamics from Microscopic Image Sequences SPIE Medical Imaging, 13-18 February 2010, San Diego, California, USA (bib) |
|
| N. Brieu, B. Glocker, N. Navab, M. Groher
MAP-MRF Optimal Partitioning for Dynamic Texture Segmentation of Thrombus in Time-Series Microscopic Images MICCAI 2010 Workshop on Spatio Temporal Image Analysis for Longitudinal and Time-Series Image Data (STIA'10), 24 September 2010, Beijing, China (bib) |
|
| D. Zikic, B. Glocker, O. Kutter, M. Groher, N. Komodakis, A. Kamen, N. Paragios, N. Navab
Linear Intensity-based Image Registration by Markov Random Fields and Discrete Optimization Medical Image Analysis, 22 April, 2010. (bib) |
|
| G.Gul-Isguder, G. Unal, M. Groher, N. Navab, A.K.Kalkan, M. Degertekin, H.Hetterich, J.Rieber
Manifold Learning for Image-Based Gating of Intravascular Ultrasound(IVUS) Pullback Sequences 5th International Workshop on Medical Imaging and Augmented Reality, Sep 2010, Beijing (bib) |
|
| M. Groher, M. Baust, D. Zikic, N. Navab
Monocular Deformable Model-to-Image Registration of Vascular Structures International Workshop on Biomedical Image Registration (WBIR), in Lübeck, Germany, July 11-13 2010. (bib) |
|
| D. Zikic, B. Glocker, O. Kutter, M. Groher, N. Komodakis, A. Kamen, N. Paragios, N. Navab
Markov Random Field Optimization for Intensity-based 2D-3D Registration SPIE Medical Imaging, San Diego, California, USA, 13-18 February 2010 (bib) |
|
| 2009 | |
| N. Brieu, J. Serbanovic-Canic, A. Cvejic, D. Stemple, W. Ouwehand
, N. Navab, M. Groher
A dynamic texture approach to semi-automatic thrombosis segmentation in in-vivo microscopic video-sequences Workshop on Microscopic Image Analysis and Application in Biology, (MIAAB 2009), Bethesda, MD (US), 3-4 September 2009 (bib) |
|
| H. Heibel, B. Glocker, M. Groher, N. Paragios, N. Komodakis, N. Navab
Discrete Tracking of Parametrized Curves IEEE Computer Society Conference on Computer Vision and Pattern Recognition (CVPR), Miami, Florida (USA), June 2009. (bib) |
|
| M. Groher, D. Zikic, N. Navab
Deformable 2D-3D Registration of Vascular Structures in a One View Scenario IEEE Trans. Med. Imag., 28 (6), pp. 847-860 (bib) |
|
| 2008 | |
| F. Bender, M. Groher, A. Khamene, W. Wein, H. Heibel, N. Navab
3D Dynamic Roadmapping for Abdominal Catheterizations Medical Image Computing and Computer-Assisted Intervention, MICCAI, 2008, New York, USA, September 6-10 2008 (bib) |
|
| C. Unger, M. Groher, N. Navab
Image Based Rendering for Motion Compensation in Angiographic Roadmapping IEEE Computer Society Conference on Computer Vision and Pattern Recognition, Anchorage, Alaska (USA), June 24-26, 2008 (bib) |
|
| D. Zikic, M. Groher, A. Khamene, N. Navab
Deformable Registration of 3D Vessel Structures to a Single Projection Image. SPIE Medical Imaging, San Diego, California, USA, 16-21 February 2008 (bib) |
|
| S. Atasoy, M. Groher, D. Zikic, B. Glocker, T. Waggershauser, M. Pfister, N. Navab
Real-Time Respiratory Motion Tracking: Roadmap Correction for Hepatic Artery Catheterizations SPIE Medical Imaging, San Diego, California, USA, 16-21 February 2008 (bib) |
|
| N. Navab, M. Groher
Novel Navigation and Advanced Visualization Techniques for Abdominal Catheterization Hospital Imaging and Radiology Europe, 2 (4), pp. 29-30, Feb. 2008 (bib) |
|
| 2007 | |
| M. Groher, F. Bender, R.T. Hoffmann, N. Navab
Segmentation-driven 2D-3D Registration for Abdominal Catheter Interventions Medical Image Computing and Computer-Assisted Intervention, MICCAI, 2007, Brisbane, Australia, October 29 - November 2 2007, LNCS 4792, pp. 527-535 (bib) |
|
| M. Groher, T. F. Jakobs, N. Padoy, N. Navab
Planning and Intraoperative Visualization of Liver Catheterizations: New CTA Protocol and 2D-3D Registration Method Academic Radiology 14 (11), pp.1324-1339. Special issue of MICCAI 2006 (bib) |
|
| L. König, M. Groher, A. Keil, Ch. Glaser, M. Reiser, N. Navab
Semi-Automatic Segmentation of the Patellar Cartilage in MRI Proc. of Bildverarbeitung für die Medizin (BVM 2007), Munich, Germany, March 2007. The original publication is available online at www.springerlink.com. (bib) |
|
| M. Groher, R.T. Hoffmann, C. J. Zech, M. Reiser, N. Navab
An Efficient Registration Algorithm for Advanced Fusion of 2D/3D Angiographic Data in Proceedings of Bildverarbeitung fuer die Medizin (BVM 2007), Munich, Germany, March 2007 (bib) |
|
| 2006 | |
| M. Groher, T. F. Jakobs, M. Reiser, N. Navab
Advanced 2D-3D Registration Method for Transarterial Chemoembolizations 5. Jahrestagung der Deutschen Gesellschaft für Computer-und Roboter-Assistierte Chirurgie (CURAC 2006), Hannover, Germany, September 2006 (bib) |
|
| M. Groher, N. Padoy, T. F. Jakobs, N. Navab
New CTA Protocol and 2D-3D Registration Method for Liver Catheterization Proceedings of Medical Image Computing and Computer-Assisted Intervention (MICCAI 2006), Copenhagen, Denmark, October 2006 (bib) |
|
| 2005 | |
| M. Groher, T. F. Jakobs, N. Padoy, N. Navab
Towards a Feature-based 2D-3D Registration Method of CTA and 2D Angiograms for Liver Tumor Chemoembolizations 4. Jahrestagung der Deutschen Gesellschaft für Computer-und Roboter-Assistierte Chirurgie (CURAC 2005), Berlin, Germany, September 2005 (bib) |
|
| 2004 | |
| R. Bauernschmitt, E.U. Schirmbeck, M. Groher, P. Keitler, M. Bauer, H. Najafi, G. Klinker, R. Lange
Navigierte Platzierung endovaskulärer Aortenstents Z Kardiologie 2004 93 S3: 116 (bib) |
|
Teaching
Finished and Currently active Diploma Theses, SEPs, IDPs I (co-)supervised
Available:- Fusion of in-vivo and ex-vivo microscopy volumes of mouse brain samples by Eleni Siampli (IDP/Klinisches Anwendungsprojekt 2012/02/01)
- Blood Flow Simulation for Angiographic Interventions by Max Hainz (DA/MA/BA 2009/06/15)
- Real-Time Respiratory Motion Tracking: Roadmap Correction for Endovascular Interventions by Selen Atasoy (DA/MA/BA 2007/06/15)
- Rapid prototyping with Open Inventor for Medical Applications by Stephan v. Watzdorf (SEP 2007/01/01)
- Hepatic Vessel Extraction for 2D/3D Registration by Nicolas Padoy (DA/MA/BA 2005/09/15)
- Development and Implementation of a User-Friendly Tool for Analysis of the Patella Cartilage Using Magnetic Resonance Tomography by Lorenz König (IDP )
- Optimization Algorithms for 3D Computer Vision: by Moritz Blume (SEP )
- Implementation of a Fast Covariant Feature Detector by Stefan Hinterstoisser (SEP )
- Segmentation, Centerline Extraction, and Graph Creation of Angiographic Vessels in 2D by Josef Minde and Gefei Fu (SEP )
- 2D / 3D Pose Estimation by Christian Waechter (SEP )
Teaching Assistance
- Lecture 3D Computer Vision II (2010WiSe)
- Lecture Medical Imaging Entrepreneurship (2010WiSe)
People I helped moving to a new flat while at the chair
3x Joerg Traub + one IKEA shopping...?Tobias Sielhorst
Pierre Georgel And I live on the fifth floor without elevator, look how nice he is
Nicolas Padoy
Ben Glocker
Olivier Pauly
It's never good to have the biggest car at a chair...
| UsersForm | |
|---|---|
| Title: | Dr. |
| Circumference of your head (in cm): | |
| Firstname: | Martin |
| Middlename: | |
| Lastname: | Groher |
| Picture: |  |
| Birthday: | |
| Nationality: | Germany |
| Languages: | |
| Groups: | Registration/Visualization, Medical Imaging |
| Expertise: | Registration/Visualization, Segmentation, Medical Imaging, Computer Vision |
| Position: | External Collaborator |
| Status: | Alumni |
| Emailbefore: | groher |
| Emailafter: | cs.tum.edu |
| Room: | microDimensions |
| Telephone: | +49 89 289 10930 |
| Alumniactivity: | CEO of microDimensions |
| Defensedate: | 8 April 2008 |
| Thesistitle: | 2D-3D Registration of Angiographic Images |
| Alumnihomepage: | |
| Personalvideo01: | |
| Personalvideotext01: | |
| Personalvideopreview01: | |
| Personalvideo02: | |
| Personalvideotext02: | |
| Personalvideopreview02: | |